- Current
- Browse
- Collections
-
For contributors
- For Authors
- Instructions to authors
- Article processing charge
- e-submission
- For Reviewers
- Instructions for reviewers
- How to become a reviewer
- Best reviewers
- For Readers
- Readership
- Subscription
- Permission guidelines
- About
- Editorial policy
Articles
- Page Path
- HOME > Diabetes Metab J > Volume 46(4); 2022 > Article
-
Original ArticleOthers Development of Various Diabetes Prediction Models Using Machine Learning Techniques
-
Juyoung Shin1,2
 , Jaewon Kim3, Chanjung Lee3, Joon Young Yoon3, Seyeon Kim3, Seungjae Song3, Hun-Sung Kim2,4
, Jaewon Kim3, Chanjung Lee3, Joon Young Yoon3, Seyeon Kim3, Seungjae Song3, Hun-Sung Kim2,4
-
Diabetes & Metabolism Journal 2022;46(4):650-657.
DOI: https://doi.org/10.4093/dmj.2021.0115
Published online: March 11, 2022
1Health Promotion Center, Seoul St. Mary’s Hospital, Seoul, Korea
2Department of Endocrinology and Metabolism, College of Medicine, The Catholic University of Korea, Seoul, Korea
3LifeSemantics Corp., Seoul, Korea
4Department of Medical Informatics, College of Medicine, The Catholic University of Korea, Seoul, Korea
-
Corresponding author: Hun-Sung Kim
 Department of Medical Informatics, College of Medicine, The Catholic University of Korea, 222 Banpo-daero, Seocho-gu, Seoul 06591, Korea E-mail: 01cadiz@hanmail.net
Department of Medical Informatics, College of Medicine, The Catholic University of Korea, 222 Banpo-daero, Seocho-gu, Seoul 06591, Korea E-mail: 01cadiz@hanmail.net
Copyright © 2022 Korean Diabetes Association
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
-
Background
- There are many models for predicting diabetes mellitus (DM), but their clinical implication remains vague. Therefore, we aimed to create various DM prediction models using easily accessible health screening test parameters.
-
Methods
- Two sets of variables were used to develop eight DM prediction models. One set comprised 62 easily accessible examination results of commonly used variables from a tertiary university hospital. The second set comprised 27 of the 62 variables included in the national routine health checkups. Gradient boosting and random forest algorithms were used to develop the models. Internal validation was performed using the stratified 10-fold cross-validation method.
-
Results
- The area under the receiver operating characteristic curve (ROC-AUC) for the 62-variable DM model making 12-month predictions for subjects without diabetes was the largest (0.928) among those of the eight DM prediction models. The ROC-AUC dropped by more than 0.04 when training with the simplified 27-variable set but still showed fairly good performance with ROC-AUCs between 0.842 and 0.880. The accuracy was up to 11.5% higher (from 0.807 to 0.714) when fasting glucose was included.
-
Conclusion
- We created easily applicable diabetes prediction models that deliver good performance using parameters commonly assessed during tertiary university hospital and national routine health checkups. We plan to perform prospective external validation, hoping that the developed DM prediction models will be widely used in clinical practice.
- Diabetes mellitus (DM) is a major chronic disease whose prevalence is continuously increasing [1]. Diabetes causes countless issues as it is associated with various complications and eventually plays a role as a major cause (direct or indirect) of death. Therefore, guidelines recommend performing early and regular screening tests for people with diabetes risk factors, including family history of diabetes, prediabetic condition, history of gestational diabetes, and insulin resistance, emphasizing the need for early diagnosis and treatment [2-4]. Furthermore, DM prevention is as important as its management. Once the risk of DM occurrence is recognized, early intervention, including healthy dietary habits and regular physical activity, helps slow down the progression from a prediabetic to a diabetic condition [5-7]. In this context, informing individuals of their risk of diabetes and offering them appropriate lifestyle modifications would be more efficient and effective than treating them once the disease has developed.
- Machine learning and data mining models, techniques in the field of artificial intelligence, have been widely used to detect meaningful variables and determine the relationship between them. Recently, these have been introduced in the medical field and used to identify hidden factors and suggest computational patterns based on large databases with various parameters [8-14]. Several models have been developed to predict diabetes under different conditions. However, their application in clinical practice has been limited as they have not provided consistent outcomes across varying datasets, designs, etc.
- Many people undergo general routine health checkups provided by the national healthcare system or according to their own needs, and huge datasets of general health checkup findings, available to the patients or for research purposes, have been accumulated over time in Korea [15]. This study aimed to develop valid and applicable DM prediction models using an electronic medical record database as a retrospective cohort study. Implementations of various DM prediction model types were attempted, taking full advantage of the available variables in our database, a dataset of private health checkups conducted at a health promotion center in a tertiary hospital. In Korea, the state requires individuals to undergo routine health examinations. However, the number of health examination variables in the tests provided by the state is much smaller than that in examinations provided by our health promotion center. Therefore, we attempted to create the same DM prediction models with a reduced variable number based on the tests performed during national routine health checkups. The reduced variable number would allow our models to be used with results from not only the relatively intensive private but also simplified national health checkups.
INTRODUCTION
- Privacy protection
- All the data obtained from the electronic medical records for this study were encrypted, and other personal information was not collected. Examinees were assigned random temporary identifications, and only anonymous data were available to observers and analysts. As this study used only anonymous data, it was impossible to violate human rights or infringe on moral or ethical issues. Therefore, it was not necessary to obtain informed consent from the participants. The Institutional Review Board of the Catholic University of Korea, Seoul St. Mary’s Hospital (IRB No. KC18RESI0708) approved the study.
- Data collection
- The data were extracted from electronic medical records of the Health Promotion Center of Seoul St. Mary’s Hospital between 2009 and 2018. We included subjects who underwent at least two full checkups. We defined diabetic patients if they fulfilled any of the following three criteria [16]: (1) self-reported diabetes; (2) taking any glucose-lowering agent; (3) fasting glucose level ≥126 mg/dL or glycosylated hemoglobin (HbA1c) ≥6.5%. Prediabetes was defined as the condition wherein the fasting glucose level is 100 to 125 mg/dL or HbA1c is 5.7% to 6.4%. Subjects in the non-diabetic group were randomly extracted from those with fasting glucose levels <126mg/dL and HbA1c <6.5% and included both normoglycemic and prediabetic subjects.
- DM prediction model types
- Four prediction models were created to ensure high accuracy (Fig. 1). Model-1 and Model-2 were models for predicting the development of DM after 2 and 1 years, respectively, in subjects without diabetes (normal or prediabetic). Model-3 was a model for predicting the development of DM after 1 year in prediabetic subjects. Model-4 was a model for predicting the development of DM after 1 year in prediabetic subjects after learning the difference between one and 2 years before diabetes diagnosis. Gradient boosting algorithms were used in Model-1, Model-2, and Model-3, and random forest algorithms were used in Model-4. The hyperparameters of each model were selected by a random search.
- Subjects with data of the previous 24 months (approximate intervals of 18 to 30 months) before DM diagnosis were included as a diabetic group in the 2-year prediction model. Similarly, subjects with data of the previous 12 months (approximate intervals of 8 to 16 months) before DM diagnosis were included as a diabetic group in the 1-year prediction model. If there were two or more visits within the same interval, the tests closest to the 12- or 24-month were used. Non-diabetic subjects were randomly selected from subjects without diabetes according to the design of each model. The number of non-diabetic and diabetic subjects was adjusted to be the same in each model.
- Variables used to develop the diabetes prediction model
- Two sets of variables were used to develop the DM prediction models. Aiming to use as many of the available variables as possible, one set included 62 easily accessible variables from commonly performed examinations at the health promotion center in the Seoul St. Mary’s hospital. To increase clinical use, another predictive model was developed using 27 of the 62 variables recorded in the national health checkups. The variables included age, sex, medication use, underlying diseases, family history, physical examinations, and laboratory results (Supplementary Table 1).
- Validation
- Using the stratified 10-fold cross-validation method, we tuned the model hyperparameters in search of the set of optimal hyperparameters that would yield the largest area under the curve (AUC) in the receiver operating characteristic (ROC) analysis. We created nine training datasets by randomly but evenly splitting 90% of subjects in the dataset and created a validation dataset with the remaining 10%. We used the training datasets to train the model and used the validation dataset to evaluate the performance of the model based on an unknown dataset. This cross-validation process was repeated 10 times, and the mean and standard deviation results were calculated (Supplementary Fig. 1).
METHODS
- Overall, the medical records contained information of approximately 134,691 individuals of which 3,952 were diagnosed with diabetes. Among these, 752 subjects with data of the previous 24 months were included in DM prediction Model-2, and 641 subjects with data of the previous 12 months were included in DM prediction Model-1. The same number of subjects (752 for Model-1 and 641 for Model-2) were randomly selected from the remaining 130,739 subjects without diabetes and assigned to the non-diabetic groups. Among the diabetic patients, 519 subjects were included in Model-3 after they were found to have prediabetic conditions. The non-diabetic group of Model-3 comprised 519 randomly selected subjects without diabetes whose laboratory results indicated that they were prediabetic 12 months before. We included 281 subjects in Model-4 after limiting those with the data of the previous 24 months (Table 1).
- Table 1 shows the performance evaluation parameters of the DM prediction models. All prediction models performed better with 62 variables than with 27 variables, with an AUC difference of over 0.04 between them. Model-2 had the largest AUCs of 0.928 and 0.880 with 62 and 27 variables, respectively. Other respective parameters of this model included accuracy (0.858 and 0.807), recall (0.856 and 0.817), and precision (0.857 and 0.802). For prediabetic subjects, Model-4 had larger AUCs (0.925 and 0.873 with 62 and 27 variables, respectively) than those of Model-3.
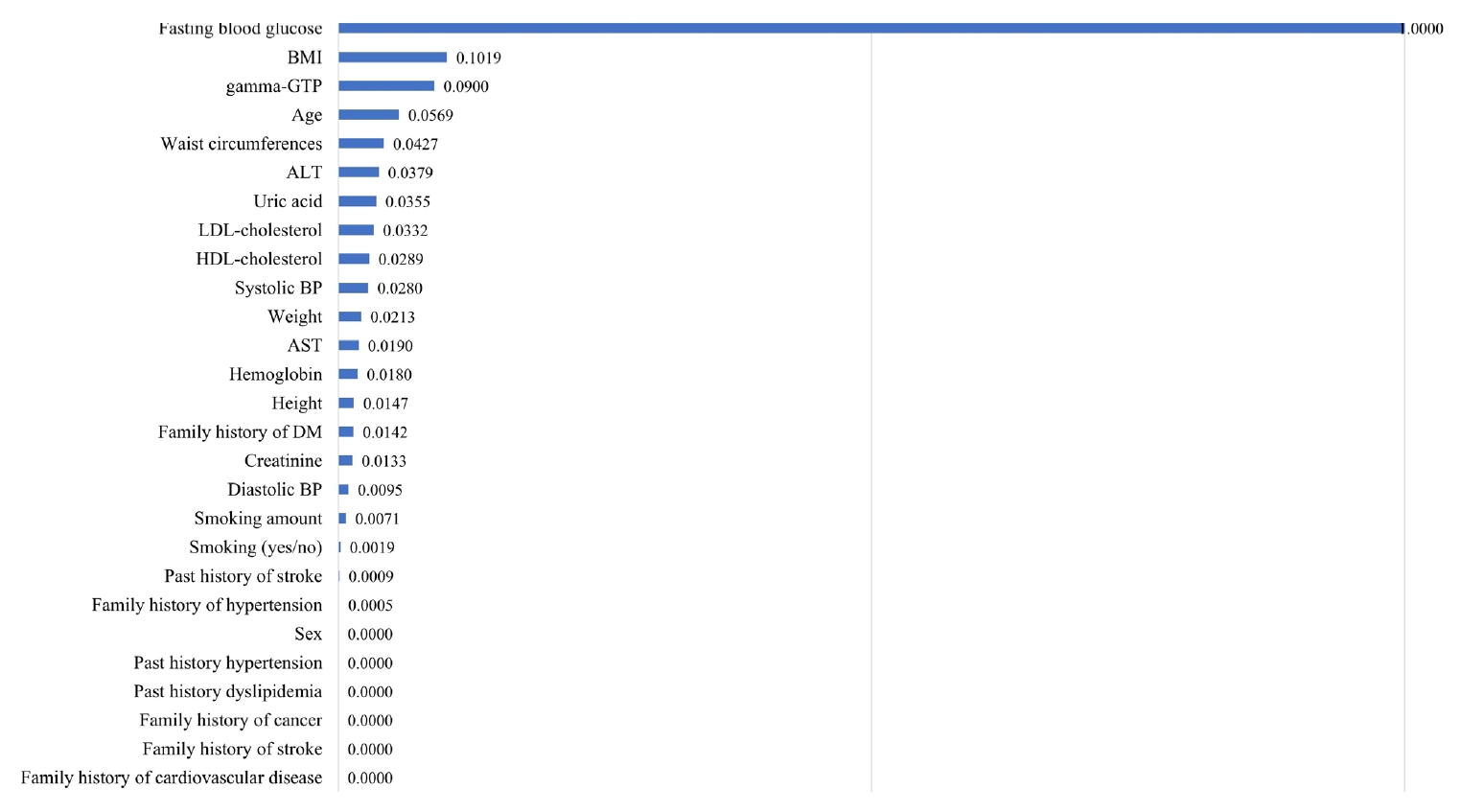
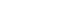
- Fig. 2 shows the effects of each of the 27 variables on diabetes prediction, listing the variables in decreasing order of importance. Fasting glucose level was the most important prediction variable, followed by body mass index (BMI). The regression coefficient and odds ratio were 0.028 and 1.028, respectively, for fasting glucose levels of –0.123 and 1.013, respectively, for BMI.
- The performance of the models changed significantly depending on whether fasting glucose was included. Table 2 shows the importance of fasting glucose in the prediction of diabetes within 2 years. After excluding the fasting glucose level (Model-1A), the accuracy decreased from 0.807 to 0.714, recall decreased from 0.817 to 0.747, precision decreased from 0.802 to 0.702, and AUC decreased from 0. 878 to 0.793. Models 1B, 1C, and 1D, which included fasting glucose, had performance parameter values similar to those of Model-1. When building a model with fasting blood glucose alone, the accuracy was 0.803 (Model-1B). Model-1E, which included the two most powerful variables, fasting blood glucose and HbA1c, had an accuracy of 0.820, a recall of 0.826, a precision of 0.818, and an AUC of 0.898. Supplementary Fig. 2 shows the distribution of the study population according to fasting glucose levels. The distribution graph of the diabetic group seems to have shifted to the right, compared to that of the non-diabetic group. The mean and standard deviation values were 107.27±9.90 mg/dL for the diabetic group and 91.36±9.79 mg/dL for the non-diabetic group. Fasting blood glucose with a cutoff value of 99 mg/dL is expected to predict the presence of diabetes within 2 years with an accuracy of 0.803.
RESULTS
- In this study, we developed 1- and 2-year diabetes prediction models using electronic medical records from general health checkups at Seoul St. Mary’s Hospital. All models showed good performance with AUCs of approximately 0.9. Silva et al. [17] suggested an AUC of 0.812 for machine learning prediction models in their systematic review and meta-analysis. In comparison, the results of this study are satisfactory.
- There are several possible reasons for the outstanding performance of our predictive models despite using a relatively small sample size. We used diverse variables including self-reported information, physical examinations, and laboratory tests. A previous study with only the information from a questionnaire showed an AUC of 0.766 despite using a larger sample size of 18,301 subjects [18]. Our models with 62 variables included pulmonary function test results. It was suggested that some of them were independent risk factors for developing diabetes, where forced vital capacity and forced expiratory volume in one-second correlated significantly with the incidence of diabetes [19]. However, to the best of our knowledge, this is the first study to present a machine learning diabetes predictive model, particularly using pulmonary function test results. Unfortunately, some meaningful variables, such as high-sensitivity C-reactive protein and adiponectin, were not included because we obtained less than 20,000 measurements/data points for these. Other meaningful variables, such as stress, occupation, and education, were not included as they were not routinely checked or recorded [20]. We also did not include information from imaging studies as they cannot be automatically entered into the computed dataset because standardized description, definite categorization, and data processing were not yet established.
- We included fasting glucose and HbA1c levels in our model. We decided to include them because of their clinical importance. The importance of the two parameters was well proven by evaluating the performance of the 2-year prediction model with and without these variables. Our results are consistent with those of a Japanese study in which the AUC increased from 0.717 to 0.893 and from 0.734 to 0.882 by adding fasting glucose and HbA1c levels, respectively [21]. Other studies also showed similar degrees of improvement when fasting glucose and HbA1c levels were included [22-27].
- The prediction time was limited to 1 or 2 years. A previous study developed a prediction model with a lower AUC of 0.820 after analyzing 93 parameters of 3,363 individuals; however, they made an eight-year prediction [20]. Generally, a longer prediction time is known to be more difficult. The performances of the 1-year prediction were well maintained with a similarly small sample size when extended to the 2-year prediction in our study. Our 2-year prediction model had a performance accuracy of over 80% when fasting blood glucose levels exceeded a certain threshold. Furthermore, the diabetic and non-diabetic groups had distinct distributions according to fasting blood glucose levels (Supplementary Fig. 2). In this context, our models could have an even better performance in predicting diabetes over a longer period such as 5 years.
- One of the study strengths was data management [28,29]. We decided to perform simple median imputation and hyperparameter tuning because little had changed in the performance with the exponential smoothing model, last observation carried forward method, Holt-Winters method, and more.
- The study population was limited to those who underwent general checkups in one center, sharing common characteristics such as homogeneous Korean ethnicity and similar socioeconomic statuses. Diabetes pathophysiology differs between ethnic groups and socioeconomic environments. For instance, the Framingham Diabetes Risk Scoring model [30], a well-known diabetes predictor based on a middle-aged United States population with an AUC of 0.850, showed different performance when applied to a Canadian population, with an AUC of 0.78 [31].
- Our goal was to reduce the economic burden on society by preventing diabetes rather than treating it. We developed models that deliver outstanding performance using 62 easily obtainable and commonly measured variables. Simplified models with 27 variables present in the national health checkups were inferior to our models with 62 variables, but their performance parameters were still similar or better than those in previous reports. As the Korean National Health Insurance Service (NHIS) provides for medical checkups every 2 years for all Koreans [15], our models could be readily applied to anyone who underwent the simple free national checkups. Individuals could recognize their risk of developing diabetes in the near future, within 1 or 2 years in this study, and individualized medical advice could be applied to prevent diabetes. Facing the specified risk could motivate people at risk to take action and emphasize a healthy lifestyle, especially for those at high-risk of diabetes. Physicians could encourage strong intervention and short-term follow-up, and politicians could devise strategies for a stratified approach to slow down the progress of DM (a society program). In this study, the model with the two most powerful variables, fasting blood glucose and HbA1c, outperformed the model with 27 variables included in NHIS checkups. Rhee et al. [32] developed a diabetes prediction model using the NHIS cohort of Korea, which outperformed the conventional model. If they can add HbA1c to their model, they might achieve outstanding performance. Therefore, we could ask the national health policymakers to add HbA1c in the routine examination for nationwide general checkups after making cost-effectiveness calculations. This will help prevent high-risk individuals from proceeding to develop diabetes as well as find undiagnosed diabetic patients, so that they can be treated. It should be worthwhile because the model with only fasting plasma glucose and HbA1c outperformed the model with the 27 national general checkup variables. We hope to determine the effectiveness of detection, followed by intervention. Fasting glucose level is important in both the diagnosis and prediction of diabetes. Kim et al. [33] suggested that higher fasting glucose levels, even within the normal range, lead to a risk of diabetes. We expect to predict the risk of diabetes in populations with normal fasting glucose levels in another study.
- This study has some limitations. Our dataset was obtained from a tertiary institution. Typically, patients with more serious and multiple diseases tend to visit higher level hospitals. However, this tendency is not expected to be too high as the data were from a screening service for diseases as general checkups and not an examination of patients, and we included a large number of examinees, especially those who underwent at least two examinations approximately 2 years apart. The examinees visited according to their own desire to be screened. They were not selected by searching for the international classification of disease codes. The model was validated internally and not externally. As we have no information except that of 62 variables, we cannot confirm that our study subjects are representative of the general population. Therefore, it would need to be validated before being applied to other situations. Typically, machine learning prediction models require validation when conditions change. It is preferable to create a specific prediction model for each location and regularly check its performance. Non-diabetic groups were randomly selected from those who met our conditions. We neither focused on the characteristics of each group nor used matching techniques to specify the weights of importance of specific variables. Instead, we put all variables together, including age and sex, and evaluated the effects of all these as real-world data.
- In conclusion, we developed a high-performance model to predict diabetes within 1 and 2 years. We expect the risk predictions suggested by the model to help health providers and clinicians provide more specific advice and encourage the examinees to lead a healthy lifestyle. We hope that this study will become a foundation for further clinical trials to elucidate the application of diabetes risk calculators, early interventions, and their diabetes prevention effects.
DISCUSSION
SUPPLEMENTARY MATERIALS
Supplementary Fig. 1.
Supplementary Fig. 2.
-
CONFLICTS OF INTEREST
The opinions expressed in this paper are those of the authors and do not necessarily represent those of LifeSemantics Corporation.
-
AUTHOR CONTRIBUTIONS
Conception or design: J.S., H.S.K.
Acquisition, analysis, or interpretation of data: J.K., C.L., J.Y.Y., S.K., S.S., and H.S.K.
Drafting the work or revising: J.S., H.S.K.
Final approval of the manuscript: H.S.K.
-
FUNDING
None
NOTES
-
Acknowledgements
- We thank Jamin Hong (LifeSemantics Corp., Seoul, Republic of Korea) for some assistance with data preprocessing.

Model-1: 27 variables; Model-1A: variables of Model-1 except fasting blood glucose; Model-1B: fasting blood glucose; Model-1C: age, sex, body mass index (BMI), fasting blood glucose; Model-1D: age, BMI, waist circumference, systolic and diastolic blood pressure, pulse rate, family history of diabetes, hemoglobin, fasting blood glucose; Model-1E: fasting blood glucose, glycosylated hemoglobin.
ROC-AUC, area under the receiver operating characteristic curve.
- 1. Kim BY, Won JC, Lee JH, Kim HS, Park JH, Ha KH, et al. Diabetes fact sheets in Korea, 2018: an appraisal of current status. Diabetes Metab J 2019;43:487-94.ArticlePubMedPMCPDF
- 2. Gillies CL, Abrams KR, Lambert PC, Cooper NJ, Sutton AJ, Hsu RT, et al. Pharmacological and lifestyle interventions to prevent or delay type 2 diabetes in people with impaired glucose tolerance: systematic review and meta-analysis. BMJ 2007;334:299.ArticlePubMedPMC
- 3. Lindstrom J, Peltonen M, Eriksson JG, Louheranta A, Fogelholm M, Uusitupa M, et al. High-fibre, low-fat diet predicts long-term weight loss and decreased type 2 diabetes risk: the Finnish Diabetes Prevention Study. Diabetologia 2006;49:912-20.ArticlePubMedPDF
- 4. Kim TM, Kim H, Jeong YJ, Baik SJ, Yang SJ, Lee SH, et al. The differences in the incidence of diabetes mellitus and prediabetes according to the type of HMG-CoA reductase inhibitors prescribed in Korean patients. Pharmacoepidemiol Drug Saf 2017;26:1156-63.ArticlePubMedPDF
- 5. Knowler WC, Barrett-Connor E, Fowler SE, Hamman RF, Lachin JM, Walker EA, et al. Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med 2002;346:393-403.ArticlePubMedPMC
- 6. Pan XR, Li GW, Hu YH, Wang JX, Yang WY, An ZX, et al. Effects of diet and exercise in preventing NIDDM in people with impaired glucose tolerance: the Da Qing IGT and Diabetes Study. Diabetes Care 1997;20:537-44.ArticlePubMedPDF
- 7. Tuomilehto J, Lindstrom J, Eriksson JG, Valle TT, Hamalainen H, Ilanne-Parikka P, et al. Prevention of type 2 diabetes mellitus by changes in lifestyle among subjects with impaired glucose tolerance. N Engl J Med 2001;344:1343-50.ArticlePubMed
- 8. Sajda P. Machine learning for detection and diagnosis of disease. Annu Rev Biomed Eng 2006;8:537-65.ArticlePubMed
- 9. Odedra D, Samanta S, Vidyarthi AS. Computational intelligence in early diabetes diagnosis: a review. Rev Diabet Stud 2010;7:252-62.ArticlePubMedPMC
- 10. Yoo I, Alafaireet P, Marinov M, Pena-Hernandez K, Gopidi R, Chang JF, et al. Data mining in healthcare and biomedicine: a survey of the literature. J Med Syst 2012;36:2431-48.ArticlePubMedPDF
- 11. Wang C, Li L, Wang L, Ping Z, Flory MT, Wang G, et al. Evaluating the risk of type 2 diabetes mellitus using artificial neural network: an effective classification approach. Diabetes Res Clin Pract 2013;100:111-8.ArticlePubMed
- 12. Barber SR, Davies MJ, Khunti K, Gray LJ. Risk assessment tools for detecting those with pre-diabetes: a systematic review. Diabetes Res Clin Pract 2014;105:1-13.ArticlePubMed
- 13. Choi SB, Kim WJ, Yoo TK, Park JS, Chung JW, Lee YH, et al. Screening for prediabetes using machine learning models. Comput Math Methods Med 2014;2014:618976.ArticlePubMedPMCPDF
- 14. Arabasadi Z, Alizadehsani R, Roshanzamir M, Moosaei H, Yarifard AA. Computer aided decision making for heart disease detection using hybrid neural network-genetic algorithm. Comput Methods Programs Biomed 2017;141:19-26.ArticlePubMed
- 15. Lee WC, Lee SY. National health screening program of Korea. J Korean Med Assoc 2010;53:363-70.Article
- 16. Kim MK, Ko SH, Kim BY, Kang ES, Noh J, Kim SK, et al. 2019 Clinical practice guidelines for type 2 diabetes mellitus in Korea. Diabetes Metab J 2019;43:398-406.ArticlePubMedPMCPDF
- 17. Silva K, Lee WK, Forbes A, Demmer RT, Barton C, Enticott J. Use and performance of machine learning models for type 2 diabetes prediction in community settings: a systematic review and meta-analysis. Int J Med Inform 2020;143:104268.ArticlePubMed
- 18. Alssema M, Vistisen D, Heymans MW, Nijpels G, Glumer C, Zimmet PZ, et al. The evaluation of screening and early detection strategies for type 2 diabetes and impaired glucose tolerance (DETECT-2) update of the Finnish diabetes risk score for prediction of incident type 2 diabetes. Diabetologia 2011;54:1004-12.ArticlePubMedPDF
- 19. Choi HS, Lee SW, Kim JT, Lee HK. The association between pulmonary functions and incident diabetes: longitudinal analysis from the Ansung Cohort in Korea. Diabetes Metab J 2020;44:699-710.ArticlePubMedPMCPDF
- 20. Casanova R, Saldana S, Simpson SL, Lacy ME, Subauste AR, Blackshear C, et al. Prediction of incident diabetes in the Jackson Heart Study using high-dimensional machine learning. PLoS One 2016;11:e0163942.ArticlePubMedPMC
- 21. Nanri A, Nakagawa T, Kuwahara K, Yamamoto S, Honda T, Okazaki H, et al. Development of risk score for predicting 3-year incidence of type 2 diabetes: Japan Epidemiology Collaboration on Occupational Health Study. PLoS One 2015;10:e0142779.ArticlePubMedPMC
- 22. Doi Y, Ninomiya T, Hata J, Hirakawa Y, Mukai N, Iwase M, et al. Two risk score models for predicting incident type 2 diabetes in Japan. Diabet Med 2012;29:107-14.ArticlePubMed
- 23. Heianza Y, Arase Y, Hsieh SD, Saito K, Tsuji H, Kodama S, et al. Development of a new scoring system for predicting the 5 year incidence of type 2 diabetes in Japan: the Toranomon Hospital Health Management Center Study 6 (TOPICS 6). Diabetologia 2012;55:3213-23.ArticlePubMedPDF
- 24. Sun F, Tao Q, Zhan S. An accurate risk score for estimation 5-year risk of type 2 diabetes based on a health screening population in Taiwan. Diabetes Res Clin Pract 2009;85:228-34.ArticlePubMed
- 25. Guasch-Ferre M, Bullo M, Costa B, Martinez-Gonzalez MA, Ibarrola-Jurado N, Estruch R, et al. A risk score to predict type 2 diabetes mellitus in an elderly Spanish Mediterranean population at high cardiovascular risk. PLoS One 2012;7:e33437.ArticlePubMedPMC
- 26. Aekplakorn W, Bunnag P, Woodward M, Sritara P, Cheepudomwit S, Yamwong S, et al. A risk score for predicting incident diabetes in the Thai population. Diabetes Care 2006;29:1872-7.ArticlePubMedPDF
- 27. Schulze MB, Weikert C, Pischon T, Bergmann MM, Al-Hasani H, Schleicher E, et al. Use of multiple metabolic and genetic markers to improve the prediction of type 2 diabetes: the EPIC-Potsdam Study. Diabetes Care 2009;32:2116-9.ArticlePubMedPMCPDF
- 28. Kim HS, Kim JH. Proceed with caution when using real world data and real world evidence. J Korean Med Sci 2019;34:e28.ArticlePubMedPMCPDF
- 29. Kim HS, Kim DJ, Yoon KH. Medical big data is not yet available: why we need realism rather than exaggeration. Endocrinol Metab (Seoul) 2019;34:349-54.ArticlePubMedPMCPDF
- 30. Wilson PW, Meigs JB, Sullivan L, Fox CS, Nathan DM, D’Agostino RB Sr. Prediction of incident diabetes mellitus in middle-aged adults: the Framingham Offspring Study. Arch Intern Med 2007;167:1068-74.ArticlePubMed
- 31. Mashayekhi M, Prescod F, Shah B, Dong L, Keshavjee K, Guergachi A. Evaluating the performance of the Framingham Diabetes Risk Scoring Model in Canadian electronic medical records. Can J Diabetes 2015;39:152-6.ArticlePubMed
- 32. Rhee SY, Sung JM, Kim S, Cho IJ, Lee SE, Chang HJ. Development and validation of a deep learning based diabetes prediction system using a nationwide population-based cohort. Diabetes Metab J 2021;45:515-25.ArticlePubMedPMCPDF
- 33. Kim MK, Han K, Koh ES, Hong OK, Baek KH, Song KH, et al. Cumulative exposure to impaired fasting glucose and future risk of type 2 diabetes mellitus. Diabetes Res Clin Pract 2021;175:108799.ArticlePubMed
REFERENCES
Figure & Data
References
Citations

- Predictive modeling for the development of diabetes mellitus using key factors in various machine learning approaches
Marenao Tanaka, Yukinori Akiyama, Kazuma Mori, Itaru Hosaka, Kenichi Kato, Keisuke Endo, Toshifumi Ogawa, Tatsuya Sato, Toru Suzuki, Toshiyuki Yano, Hirofumi Ohnishi, Nagisa Hanawa, Masato Furuhashi
Diabetes Epidemiology and Management.2024; 13: 100191. CrossRef - Validation of the Framingham Diabetes Risk Model Using Community-Based KoGES Data
Hye Ah Lee, Hyesook Park, Young Sun Hong
Journal of Korean Medical Science.2024;[Epub] CrossRef - Integrated Embedded system for detecting diabetes mellitus using various machine learning techniques
Rishita Konda, Anuraag Ramineni, Jayashree J, Niharika Singavajhala, Sai Akshaj Vanka
EAI Endorsed Transactions on Pervasive Health and Technology.2024;[Epub] CrossRef - The Present and Future of Artificial Intelligence-Based Medical Image in Diabetes Mellitus: Focus on Analytical Methods and Limitations of Clinical Use
Ji-Won Chun, Hun-Sung Kim
Journal of Korean Medical Science.2023;[Epub] CrossRef - Machine learning for predicting diabetic metabolism in the Indian population using polar metabolomic and lipidomic features
Nikita Jain, Bhaumik Patel, Manjesh Hanawal, Anurag R. Lila, Saba Memon, Tushar Bandgar, Ashutosh Kumar
Metabolomics.2023;[Epub] CrossRef - Retrospective cohort analysis comparing changes in blood glucose level and body composition according to changes in thyroid‐stimulating hormone level
Hyunah Kim, Da Young Jung, Seung‐Hwan Lee, Jae‐Hyoung Cho, Hyeon Woo Yim, Hun‐Sung Kim
Journal of Diabetes.2022; 14(9): 620. CrossRef - Improving Machine Learning Diabetes Prediction Models for the Utmost Clinical Effectiveness
Juyoung Shin, Joonyub Lee, Taehoon Ko, Kanghyuck Lee, Yera Choi, Hun-Sung Kim
Journal of Personalized Medicine.2022; 12(11): 1899. CrossRef

 KDA
KDA PubReader
PubReader ePub Link
ePub Link Cite
Cite