- Current
- Browse
- Collections
-
For contributors
- For Authors
- Instructions to authors
- Article processing charge
- e-submission
- For Reviewers
- Instructions for reviewers
- How to become a reviewer
- Best reviewers
- For Readers
- Readership
- Subscription
- Permission guidelines
- About
- Editorial policy
Articles
- Page Path
- HOME > Diabetes Metab J > Volume 43(5); 2019 > Article
-
Original ArticlePathophysiology Essential Role of Protein Arginine Methyltransferase 1 in Pancreas Development by Regulating Protein Stability of Neurogenin 3
-
Kanghoon Lee1
 , Hyunki Kim1
, Hyunki Kim1 , Joonyub Lee1, Chang-Myung Oh1,2, Heein Song1, Hyeongseok Kim1, Seung-Hoi Koo3, Junguee Lee4, Ajin Lim1, Hail Kim1,5
, Joonyub Lee1, Chang-Myung Oh1,2, Heein Song1, Hyeongseok Kim1, Seung-Hoi Koo3, Junguee Lee4, Ajin Lim1, Hail Kim1,5
-
Diabetes & Metabolism Journal 2019;43(5):649-658.
DOI: https://doi.org/10.4093/dmj.2018.0232
Published online: April 8, 2019
1Graduate School of Medical Science and Engineering, Korea Advanced Institute of Science and Technology, Daejeon, Korea.
2Department of Internal Medicine, CHA Bundang Medical Center, CHA University, Seongnam, Korea.
3Division of Life Sciences, Korea University, Seoul, Korea.
4Department of Pathology, Daejeon St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Daejeon, Korea.
5KAIST Institute for the BioCentury, Korea Advanced Institute of Science and Technology, Daejeon, Korea.
- Corresponding author: Hail Kim. Graduate School of Medical Science and Engineering, Korea Advanced Institute of Science and Technology, 291 Daehak-ro, Yuseong-gu, Daejeon 34141, Korea. hailkim@kaist.edu
- *Kanghoon Lee and Hyunki Kim contributed equally to this study as first authors.
Copyright © 2019 Korean Diabetes Association
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
-
Background
- Protein arginine methyltransferase 1 (PRMT1) is a major enzyme responsible for the formation of methylarginine in mammalian cells. Recent studies have revealed that PRMT1 plays important roles in the development of various tissues. However, its role in pancreas development has not yet been elucidated.
-
Methods
- Pancreatic progenitor cell-specific Prmt1 knock-out (Prmt1 PKO) mice were generated and characterized for their metabolic and histological phenotypes and their levels of Neurog3 gene expression and neurogenin 3 (NGN3) protein expression. Protein degradation assays were performed in mPAC cells.
-
Results
- Prmt1 PKO mice showed growth retardation and a severely diabetic phenotype. The pancreatic size and β-cell mass were significantly reduced in Prmt1 PKO mice. Proliferation of progenitor cells during the secondary transition was decreased and endocrine cell differentiation was impaired. These defects in pancreas development could be attributed to the sustained expression of NGN3 in progenitor cells. Protein degradation assays in mPAC cells revealed that PRMT1 was required for the rapid degradation of NGN3.
-
Conclusion
- PRMT1 critically contributes to pancreas development by destabilizing the NGN3 protein.
- Protein arginine methylation, which is one of the most common post-translational modifications of proteins in mammalian cells [1], is mediated by the enzymes of the protein arginine methyltransferase (PRMT) family [2]. By methylating various protein substrates, PRMT family members play diverse roles in gene expression, RNA splicing, signal transduction and other processes [3456]. Among the nine isoforms of PRMT, PRMT1 is the major isoform and accounts for approximately 85% of methylarginines in mammalian cells [7]. PRMT1 is ubiquitously expressed in most cells and, unsurprisingly, Prmt1 whole body knock-out (KO) mice are embryonically lethal [89]. Recently, cell type-specific functions of PRMT1 have been investigated using conditional KO mouse models. For example, deletion of Prmt1 from neural progenitors using Nestin-Cre resulted in brain demyelination due to defective development of oligodendrocytes [10], while deletion of Prmt1 from myogenic precursors using Pax7-CreERT2 resulted in the failure of muscle differentiation [11]. Thus, PRMT1 is thought play important roles in the development of different organs.
- The pancreas is the central organ responsible for regulating whole-body glucose homeostasis and metabolism by secreting diverse hormones and digestive enzymes [12]. Therefore, proper development of pancreatic endocrine and exocrine cells is essential for the maintenance of normal glycemic control. In the mouse, pancreatic specification initiates at embryonic day 8.5 (E8.5) from pancreatic and duodenal homeobox 1 (PDX1)-positive pancreatic progenitor cells [13]. At E9.5, pancreatic budding and branching morphogenesis occur. At E14.5, the so-called secondary transition occurs, during which pancreatic progenitor cells rapidly proliferate and differentiate [14]. After E14.5, individual pancreatic lineages undergo further differentiation, expansion, and organization [14]. However, although many studies have examined pancreas development, the precise regulatory mechanisms remain unknown. Here, we aimed to study the role of PRMT1 in pancreas development.
INTRODUCTION
- Mouse experiments
- Prmt1 floxed (Prmt1fl/fl) (MGI: 4432476) [15] mice were crossed with Pdx1-Creearly (MGI: 2684317) [16] mice to generate pancreatic progenitor cell-specific Prmt1 knock-out (Pdx1-Creearly; Prmt1fl/fl, herein called Prmt1 PKO) mice. Mice were housed in climate-controlled, specific pathogen-free barrier facilities under a 12-hour light/dark cycle, and chow and water were provided ad libitum. Noon on the morning of vaginal plug discovery was considered E0.5. The animal experiment protocols for this study were approved by the Institutional Animal Care and Use Committee of the Korea Advanced Institute of Science and Technology (KA2011-29). All experiments were performed in accordance with the relevant guidelines and regulations. Body weight and random blood glucose levels of mice were measured in the daytime, with the latter assessed using a Gluco Dr. Plus glucometer (Allmedicus, Anyang, Korea).
- Pancreatic tissue preparation
- The pancreata of mice were fixed with 4% paraformaldehyde for 2 to 4 hours at 4℃ and washed for 30 minutes with phosphate buffered saline (PBS) at 4℃. Pancreatic tissues were processed with an automatic tissue processor (TP1020; Leica Biosystems, Wetzlar, Germany) and embedded in molten paraffin wax. Paraffin-embedded tissue sections were sliced at a thickness of 4 µm and mounted on adhesive glass slides (081000; Marienfeld, Lauda-Königshofen, Germany).
- Hematoxylin and eosin staining
- Formalin-fixed paraffin-embedded pancreatic tissue slides were deparaffinized and rehydrated. Hematoxylin and eosin staining was performed as previously described with slight modification [17]. Images were acquired using a bright-field microscope (DS-Ri2 camera; Nikon, Tokyo, Japan) and analyses were performed with the NIS-Elements BR (Nikon) software.
- Immunofluorescence staining
- Formalin-fixed paraffin-embedded pancreatic tissue slides were deparaffinized and rehydrated. Antigen retrieval was performed by incubating the slides in sodium citrate buffer (10 mM sodium citrate, pH 6.0) for 20 minutes at 95℃. The slides were cooled for 10 minutes at room temperature and washed in PBS for 10 minutes, and the samples were blocked with 4% normal goat serum (005-000-121; Jackson ImmunoResearch, West Grove, PA, USA) in PBS for 1 hour at room temperature. The samples were then incubated for 18 hours at 4℃ with primary antibodies against the following: insulin (A0564, 1:1,000; Dako, Carpinteria, CA, USA), PRMT1 (84361, 1:1,000; Abcam, Cambridge, MA, USA), glucagon (G2654, 1:1,000; Sigma-Aldrich, St. Louis, MO, USA), amylase (A8273, 1:200; Sigma-Aldrich,), mucin 1 (MUC1, MA5-11202, 1:1,000; Invitrogen, Carlsbad, CA, USA), PDX1 (F6A11, 1:500, Developmental Studies Hybridoma Bank [DSHB], Iowa City, IA, USA; 47267, 1:1,000, Abcam), SRY-box 9 (SOX9, AB5535, 1:1,000; Merck Millipore, Burlington, MA, USA), neurogenin 3 (NGN3, F25A1B3, 1:500; DSHB), NK2 homeobox 2 (NKX2.2, 74.5A5, 1:500; DSHB), NKX6.1 (F65A2, 1:500; DSHB), ISL LIM homeobox 1 (ISL1, 40.2D6, 1:500; DSHB), and phosphorylated histone H3 (06-570, Merck Millipore, 1:1,000). The samples were washed in PBS for 10 minutes and incubated for 2 hours at room temperature with the following secondary antibodies: Alexa Fluor 647-conjugated anti-hamster immunoglobulin G (IgG, 127-605-160, 1:1,000; Jackson ImmunoResearch), Alexa Fluor 647-conjugated anti-guinea pig IgG (106-605-003, 1:1,000; Jackson ImmunoResearch), Alexa Fluor 488-conjugated anti-guinea pig IgG (106-545-003, 1:1,000; Jackson ImmunoResearch), Alexa Fluor 488-conjugated anti-rabbit IgG (111-545-144, 1:1,000; Jackson ImmunoResearch), Alexa Fluor 594-conjugated anti-rabbit IgG (111-585-144, 1:1,000; Jackson ImmunoResearch), Alexa Fluor 488-conjugated anti-mouse IgG (115-545-166, 1:1,000; Jackson ImmunoResearch), and Alexa Fluor 594-conjugated anti-mouse IgG (115-585-166, 1:1,000; Jackson ImmunoResearch). The samples were washed in PBS for 10 minutes, incubated for 5 minutes with 4′,6-diamidino-2-phenylindole (DAPI, D9542, 1 µg/mL; Sigma-Aldrich) at room temperature, and then mounted with fluorescence mounting medium (S3023; Dako). Images were acquired using a fluorescence microscope (DS-Ri2 camera; Nikon) and a confocal microscope (LSM 780; Carl Zeiss, Oberkochen, Germany). Imaging analyses were performed using the NIS-Elements BR (Nikon) and ZEN (Carl Zeiss) software packages.
- Quantitative reverse transcription polymerase chain reaction
- Total RNA was extracted from dorsal pancreas tissues using the TRIzol reagent (15596026; Invitrogen) according to the manufacturer's protocol. Genomic DNA was removed using a TURBO DNA-free kit (AM1907; Invitrogen) and 1 µg of total RNA was used to generate complementary DNA (cDNA) with a High-Capacity cDNA Reverse Transcription kit (4368813; Applied Biosystems, Waltham, MA, USA). Quantitative reverse transcription polymerase chain reaction (qRT-PCR) was performed with the Fast SYBR Green Master Mix (4385614; Applied Biosystems) and a Viia 7 Real-time PCR System (Applied Biosystems) according to the manufacturer's instructions. Relative Neurog3 expression was analyzed using the delta Ct (threshold cycle) method [18], with the Rplp0 (36B4) detected as a reference gene. The following primers were used to analyze gene expression levels: Rplp0 forward (GAGGAATCAGATGAGGATATGGGA), Rplp0 reverse (AAGCAGGCTGACTTGGT-TGC), Neurog3 forward (CAGTCACCCACTT-CTGCTTC), and Neurog3 reverse (GAGTCGGGAGAACTAGGATG).
- Immunoblotting
- Dorsal pancreas tissues obtained from mouse embryos at E14.5 were lysed with lysis buffer (10 mM Tris-Cl, 66 mM EDTA, 150 mM NaCl, 0.4% sodium deoxycholate, and 1% NP-40) for 1 hour. The samples were boiled at 100℃ for 5 minutes in sample buffer (100 mM Tris-Cl, 4% SDS, 0.2% bromophenol blue, 20% glycerol, and 200 mM β-mercaptoethanol), centrifuged for 5 minutes at 5,900 G, separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto a polyvinylidene fluoride membrane (Merck Millipore). The membrane was blocked with 5% dry milk, 0.1% Tween-20 in PBS for 1 hour and blotted with primary antibodies against PRMT1 (84361, 1:1,000; Abcam), NGN3 (F25A1B3, 1:1,000; DSHB) and α-tubulin (sc-8035, 1:1,000; Santa Cruz Biotechnology, Santa Cruz, CA, USA) in blocking solution for 18 hours at 4℃. The blot was then incubated with horseradish peroxidase-conjugated secondary antibodies (Cell Signaling Technology, Danvers, MA, USA) for 1 hour, and proteins were visualized using a ChemiDoc System (Bio-Rad, Hercules, CA, USA).
- Protein degradation assay
- Adenoviruses expressing nonspecific RNAi or Prmt1 RNAi [15] were transduced into cells 24 hours before transfection. Hemagglutinin (HA)-NGN3 plasmids were transfected into 30% to 50% confluent mPAC cells in a 6-well plate using cationic lipids (FuGene 6; Roche, Basel, Switzerland) according to the manufacturer's protocol. Additional protein synthesis was blocked 48 hours after transfection by the addition of cycloheximide (CHX) at 100 µg/mL. Cells were lysed with RIPA buffer at 0, 7.5, 15, 30, 60, and 90 minutes after CHX treatment.
- Quantification analysis
- Cell counting and β-cell area measurements were done using the ImageJ (National Institutes of Health) program. For quantitative analysis, pancreata of age-matched wild type (WT) littermates and Prmt1 PKO mice were whole-sectioned at 4 µm. Every fifth (20 µm apart) sections was selected for immunofluorescence staining. The β-cell area was calculated by dividing the insulin-positive area by the total pancreatic area.
- Statistical analysis
- All values are expressed as the mean±standard error of mean. The two-tailed Student's t-test or one-way analysis of variance (ANOVA) followed by post hoc Bonferroni's test were used to compare groups. P values below 0.05 were considered statistically significant. The levels of significance indicated in the graphs are P<0.05, P<0.01, and P<0.001.
METHODS
- Defective pancreas development in Prmt1 PKO mice
- To explore the functional role of PRMT1 in pancreas development, we generated Prmt1 PKO mice by crossing Prmt1fl/fl mice with Pdx1-Creearly mice. Prmt1 PKO was confirmed by immunofluorescence staining in embryonic pancreas at E10.5, which showed that there was no detectable level of PRMT1 in PDX1-positive pancreatic progenitor cells (Fig. 1A). Prmt1 PKO mice were born normally at the expected Mendelian ratio. However, Prmt1 PKO mice exhibited growth retardation, a lower body weight and severe hyperglycemia, with a blood glucose level greater than 500 mg/dL at 4 weeks of age (Fig. 1B–D). In contrast, heterozygous Prmt1 PKO (Pdx1-Creearly; Prmt1fl/+) mice were normal in their body weight and blood glucose level.
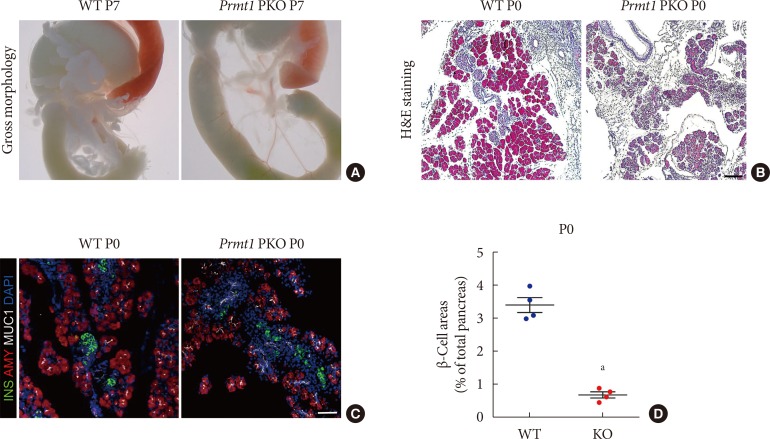
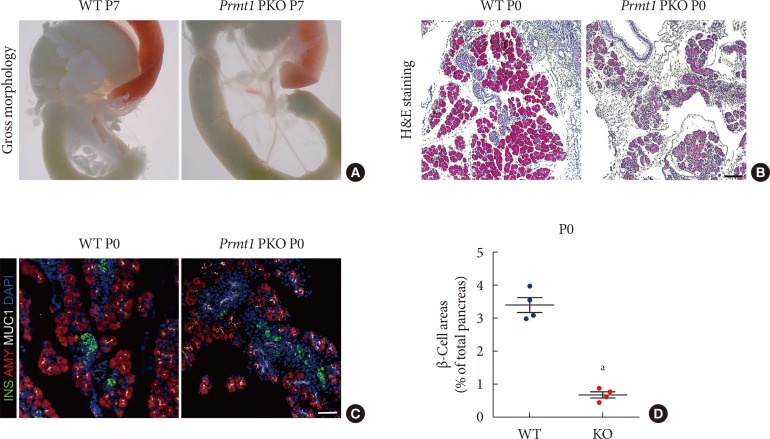
- Gross morphological analysis at postnatal day 7 (P7) showed that Prmt1 PKO mice were characterized by a severely hypoplastic pancreas (Fig. 2A). Further histological analysis with H&E staining revealed that the numbers of acini, ductal cells and islet cells were robustly reduced in Prmt1 PKO pancreas at P0, and that the Prmt1 PKO pancreas was mostly composed of progenitor epithelial cells (Fig. 2B). Immunofluorescence staining using anti-insulin and anti-amylase antibodies revealed that the numbers of acinar cells and β-cells were reduced in Prmt1 PKO pancreas (Fig. 2C). Further quantitative analysis confirmed that the β-cell area was significantly reduced in the pancreata of Prmt1 PKO mice (Fig. 2D). In addition, co-immunostaining for the ductal epithelial cell marker, MUC1, plus insulin showed that the progenitor epithelial plexus, which normally disappears during the late embryonic period, was still evident in the Prmt1 PKO pancreas at P0 (Fig. 2C). Thus, pancreas development, including both exocrine and endocrine cells, was severely compromised in Prmt1 PKO mice. These data indicate that PRMT1 is required for the normal development of the pancreas.
- PRMT1 is required for endocrine cell commitment during the secondary transition
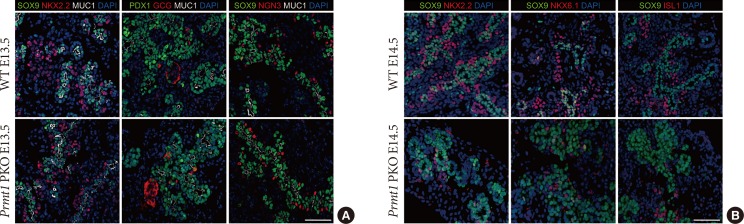
- To investigate the underlying cause of the hypoplastic pancreas seen in Prmt1 PKO mice, we carefully observed their pancreas development. Until E13.5, Prmt1 PKO mice showed normal pancreas development, with normal expression of pancreas development markers, such as NKX2.2, PDX1, SOX9, and NGN3 (Fig. 3A). However, at E14.5, severe defects in endocrine development were observed in Prmt1 PKO mice, with robust decreases in the numbers of cells that expressed the endocrine cell differentiation markers, NKX2.2, NKX6.1, and ISL1 (Fig. 3B). These data indicate that PRMT1 plays an important role in endocrine cell differentiation during the secondary transition.
- Prmt1 PKO mice exhibit increased numbers of NGN3-expressing cells during the secondary transition
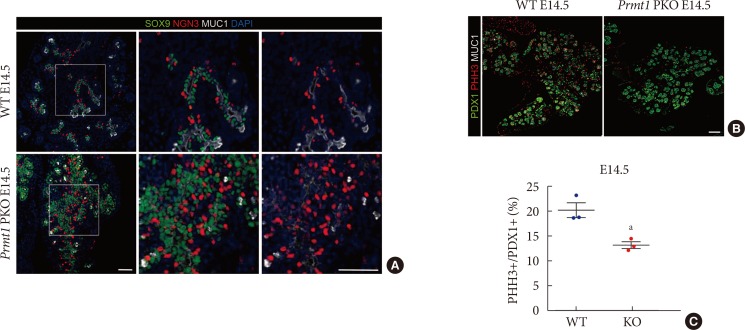
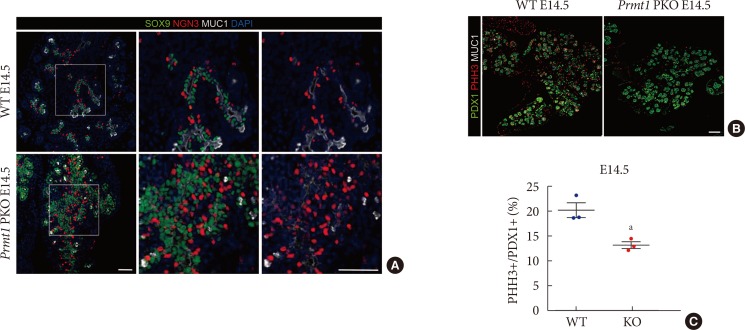
- The secondary transition, which is a critical step in pancreas development, is characterized by neogenesis and differentiation of endocrine cells, as well as active proliferation of progenitor cells [19]. During this stage, transient expression of NGN3 plays a central role in regulating the formation of endocrine cells [202122]. Sustained expression of NGN3 in pancreatic progenitor cells results in premature endocrine cell differentiation and defects in exocrine cell development [23]. These previous data motivated us to test NGN3 expression in Prmt1 PKO mice. Notably, we found that Prmt1 PKO mice showed a robust increase in the number of NGN3-positive cells that remained in the SOX9-positive trunk epithelium at E14.5 (Fig. 4A). Immunofluorescence staining of NGN3, SOX9, and MUC1 indicated that the NGN3-positive cells in Prmt1 PKO mice failed to delaminate and further differentiate into endocrine cells (Fig. 4A). During the secondary transition, NGN3-positive cells are well known to upregulate the Notch signaling of surrounding cells to induce proliferation [24]. Interestingly, co-immunostaining of the proliferation marker, phosphorylated histone H3, plus PDX1 showed that the proliferation of pancreatic progenitors was significantly reduced in Prmt1 PKO mice compared to WT mice (Fig. 4B and C). These data indicate that PRMT1 is necessary for proper NGN3 expression in pancreatic cells during the secondary transition, which is crucial for normal development of the pancreas.
- PRMT1 is necessary for rapid degradation of the NGN3 protein
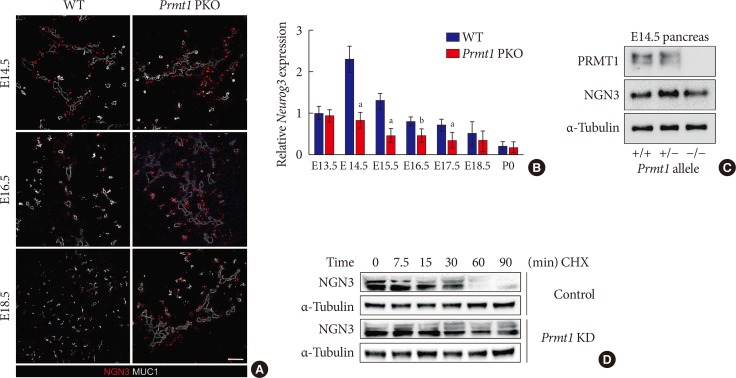
- During pancreas development in the mouse, NGN3 expression peaks at E14.5-15.5 and rapidly decreases within 24 hours [25]. However, we observed prolonged expression of NGN3 until E18.5 in the pancreas of Prmt1 PKO mice (Fig. 5A). To investigate the mechanism underlying this prolonged expression of NGN3 in the pancreas of Prmt1 PKO mice, we examined Neurog3 gene expression by qRT-PCR and NGN3 protein expression by Western blot analysis. As the mRNA and protein expressions of NGN3 were unaltered in the pancreas of Prmt1 PKO mice (Fig. 5B and C), we hypothesized that PRMT1 may be involved in altering the stability of the NGN3 protein. To test this hypothesis, we performed protein degradation assays in mPAC cells exposed to CHX, which blocks the translational elongation step of protein synthesis. Interestingly, knock-down of Prmt1 in mPAC cells was found to trigger defects in NGN3 protein degradation (Fig. 5D). These results indicate that PRMT1 is required for the rapid degradation of the NGN3 protein, but does not alter its protein expression.
RESULTS
- In this study, we demonstrated the functional role of PRMT1 in pancreas development. Prmt1 PKO mice showed severe hyperglycemia with a hypoplastic pancreas. In these mice, developmental defects of the pancreas were observed beginning at the secondary transition. Notably, NGN3-expressing cells were robustly increased in the pancreas of Prmt1 PKO mice from E14.5, and this was due to failure of the rapid degradation of NGN3 that was seen in WT mice. Indeed, our results confirmed that PRMT1 is necessary for rapid degradation of the NGN3 protein.
- Pancreas development comprises sophisticated cascades of transcription factor expressions that directs the differentiation and proliferation of individual pancreatic cell types [19]. Here, we show that PRMT1 is required for the transient expression of NGN3 during the secondary transition. However, it is still unclear how the observed increase in NGN3-positive cells translates to the severe hypoplastic phenotype of the pancreas in Prmt1 PKO mice. Based on the expression level and spatial localization of NGN3, individual NGN3-positive cells can differ in their developmental fate and lineage commitment [2627]. Therefore, precise time course-observations of NGN3-positive cells by lineage-tracing experiments are necessary in Prmt1 PKO mice.
- Further studies are also needed to uncover the detailed mechanism through which PRMT1 regulates NGN3 protein stability. Recent work has shown that cyclin dependent kinase (CDK) induces the multiple phosphorylation of NGN3, thereby promoting its protein degradation via the ubiquitin-proteasome pathway [2829]. Therefore, PRMT1 may destabilize the NGN3 protein either directly via methylation or indirectly via enhancement of phosphorylation.
- Since PRMT1 has a broad substrate specificity, it may play NGN3-independent roles in pancreas development may exist. Through its well-known function as a transcriptional coactivator, PRMT1 potentiates gene expression levels by recruitment of coactivator associated arginine methyltransferase 1 (CARM1) (PRMT4) or methylation of histone H4 [330]. In this manner, PRMT1 can directly turn on gene subsets that are related to pancreatic endocrine and exocrine cell development. On the other hand, PRMT1 modifies the activities of signaling proteins via direct methylation to regulate pathways that are essential for tissue development, such as the Wnt/β-catenin, transforming growth factor β and Notch signaling pathways [53132]. Collectively, these effects plus the up-regulation of NGN3-expressing cells may largely account for the phenotype of Prmt1 PKO mice.
- In conclusion, we herein show that PRMT1 plays an essential role in pancreas development by regulating NGN3 protein stability. Our work provides a novel mechanistic insight into pancreas development and may inform islet regeneration studies, and thus has potential implications for the treatment of diabetes.
DISCUSSION
-
Acknowledgements
- We thank Hee-Saeng Jung for technical advice and support. This work was supported by grants from the National Research Foundation (NRF) funded by the Ministry of Science and ICT, Republic of Korea (Grant numbers: NRF-2018R1A6A3A01012333 to Hyeongseok Kim, NRF-2016R1D1A1B04931995 to Ajin Lim, NRF-2013M3A9D5072550 and NRF-2015M3A9B3028218 to Hail Kim) and the KAIST Institute for the BioCentury (Grant number: N10180027 to Hail Kim).
ACKNOWLEDGMENTS
-
CONFLICTS OF INTEREST: No potential conflict of interest relevant to this article was reported.
-
AUTHOR CONTRIBUTIONS:
NOTES
- 1. Bedford MT, Clarke SG. Protein arginine methylation in mammals: who, what, and why. Mol Cell 2009;33:1-13. ArticlePubMedPMC
- 2. Wei H, Mundade R, Lange KC, Lu T. Protein arginine methylation of non-histone proteins and its role in diseases. Cell Cycle 2014;13:32-41. ArticlePubMed
- 3. Kleinschmidt MA, Streubel G, Samans B, Krause M, Bauer UM. The protein arginine methyltransferases CARM1 and PRMT1 cooperate in gene regulation. Nucleic Acids Res 2008;36:3202-3213. ArticlePubMedPMC
- 4. Ohkura N, Takahashi M, Yaguchi H, Nagamura Y, Tsukada T. Coactivator-associated arginine methyltransferase 1, CARM1, affects pre-mRNA splicing in an isoform-specific manner. J Biol Chem 2005;280:28927-28935. ArticlePubMed
- 5. Cha B, Kim W, Kim YK, Hwang BN, Park SY, Yoon JW, Park WS, Cho JW, Bedford MT, Jho EH. Methylation by protein arginine methyltransferase 1 increases stability of Axin, a negative regulator of Wnt signaling. Oncogene 2011;30:2379-2389. ArticlePubMedPDF
- 6. Blanc RS, Richard S. Arginine methylation: the coming of age. Mol Cell 2017;65:8-24. ArticlePubMed
- 7. Tang J, Frankel A, Cook RJ, Kim S, Paik WK, Williams KR, Clarke S, Herschman HR. PRMT1 is the predominant type I protein arginine methyltransferase in mammalian cells. J Biol Chem 2000;275:7723-7730. ArticlePubMed
- 8. Wada K, Inoue K, Hagiwara M. Identification of methylated proteins by protein arginine N-methyltransferase 1, PRMT1, with a new expression cloning strategy. Biochim Biophys Acta 2002;1591:1-10. ArticlePubMed
- 9. Pawlak MR, Scherer CA, Chen J, Roshon MJ, Ruley HE. Arginine N-methyltransferase 1 is required for early postimplantation mouse development, but cells deficient in the enzyme are viable. Mol Cell Biol 2000;20:4859-4869. ArticlePubMedPMCPDF
- 10. Hashimoto M, Murata K, Ishida J, Kanou A, Kasuya Y, Fukamizu A. Severe hypomyelination and developmental defects are caused in mice lacking protein arginine methyltransferase 1 (PRMT1) in the central nervous system. J Biol Chem 2016;291:2237-2245. ArticlePubMed
- 11. Blanc RS, Vogel G, Li X, Yu Z, Li S, Richard S. Arginine methylation by PRMT1 regulates muscle stem cell fate. Mol Cell Biol 2017;37:e00457-16. ArticlePubMedPMCPDF
- 12. Roder PV, Wu B, Liu Y, Han W. Pancreatic regulation of glucose homeostasis. Exp Mol Med 2016;48:e219. ArticlePubMedPMCPDF
- 13. Burlison JS, Long Q, Fujitani Y, Wright CV, Magnuson MA. Pdx-1 and Ptf1a concurrently determine fate specification of pancreatic multipotent progenitor cells. Dev Biol 2008;316:74-86. ArticlePubMedPMC
- 14. Oliver-Krasinski JM, Stoffers DA. On the origin of the beta cell. Genes Dev 2008;22:1998-2021. ArticlePubMedPMC
- 15. Choi D, Oh KJ, Han HS, Yoon YS, Jung CY, Kim ST, Koo SH. Protein arginine methyltransferase 1 regulates hepatic glucose production in a FoxO1-dependent manner. Hepatology 2012;56:1546-1556. ArticlePubMed
- 16. Gu G, Dubauskaite J, Melton DA. Direct evidence for the pancreatic lineage: NGN3+ cells are islet progenitors and are distinct from duct progenitors. Development 2002;129:2447-2457. ArticlePubMedPDF
- 17. Fischer AH, Jacobson KA, Rose J, Zeller R. Hematoxylin and eosin staining of tissue and cell sections. CSH Protoc 2008;2008:pdb.prot4986. ArticlePubMed
- 18. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001;25:402-408. ArticlePubMed
- 19. Pan FC, Wright C. Pancreas organogenesis: from bud to plexus to gland. Dev Dyn 2011;240:530-565. ArticlePubMed
- 20. Gradwohl G, Dierich A, LeMeur M, Guillemot F. Neurogenin3 is required for the development of the four endocrine cell lineages of the pancreas. Proc Natl Acad Sci U S A 2000;97:1607-1611. ArticlePubMedPMC
- 21. Lee JC, Smith SB, Watada H, Lin J, Scheel D, Wang J, Mirmira RG, German MS. Regulation of the pancreatic pro-endocrine gene neurogenin3. Diabetes 2001;50:928-936. ArticlePubMedPDF
- 22. Johansson KA, Dursun U, Jordan N, Gu G, Beermann F, Gradwohl G, Grapin-Botton A. Temporal control of neurogenin3 activity in pancreas progenitors reveals competence windows for the generation of different endocrine cell types. Dev Cell 2007;12:457-465. ArticlePubMed
- 23. Apelqvist A, Li H, Sommer L, Beatus P, Anderson DJ, Honjo T, Hrabe de Angelis M, Lendahl U, Edlund H. Notch signalling controls pancreatic cell differentiation. Nature 1999;400:877-881. ArticlePubMedPDF
- 24. Bankaitis ED, Bechard ME, Wright CV. Feedback control of growth, differentiation, and morphogenesis of pancreatic endocrine progenitors in an epithelial plexus niche. Genes Dev 2015;29:2203-2216. ArticlePubMedPMC
- 25. Rukstalis JM, Habener JF. Neurogenin3: a master regulator of pancreatic islet differentiation and regeneration. Islets 2009;1:177-184. ArticlePubMed
- 26. Bechard ME, Bankaitis ED, Hipkens SB, Ustione A, Piston DW, Yang YP, Magnuson MA, Wright CV. Precommitment low-level Neurog3 expression defines a long-lived mitotic endocrine-biased progenitor pool that drives production of endocrine-committed cells. Genes Dev 2016;30:1852-1865. ArticlePubMedPMC
- 27. Yu XX, Qiu WL, Yang L, Li LC, Zhang YW, Xu CR. Dynamics of chromatin marks and the role of JMJD3 during pancreatic endocrine cell fate commitment. Development 2018;145:dev163162. ArticlePubMedPDF
- 28. Krentz NAJ, van Hoof D, Li Z, Watanabe A, Tang M, Nian C, German MS, Lynn FC. Phosphorylation of NEUROG3 links endocrine differentiation to the cell cycle in pancreatic progenitors. Dev Cell 2017;41:129-142. ArticlePubMedPMC
- 29. Azzarelli R, Hurley C, Sznurkowska MK, Rulands S, Hardwick L, Gamper I, Ali F, McCracken L, Hindley C, McDuff F, Nestorowa S, Kemp R, Jones K, Gottgens B, Huch M, Evan G, Simons BD, Winton D, Philpott A. Multi-site neurogenin3 phosphorylation controls pancreatic endocrine differentiation. Dev Cell 2017;41:274-286. ArticlePubMedPMC
- 30. Huang S, Litt M, Felsenfeld G. Methylation of histone H4 by arginine methyltransferase PRMT1 is essential in vivo for many subsequent histone modifications. Genes Dev 2005;19:1885-1893. ArticlePubMedPMC
- 31. Xu J, Wang AH, Oses-Prieto J, Makhijani K, Katsuno Y, Pei M, Yan L, Zheng YG, Burlingame A, Bruckner K, Derynck R. Arginine methylation initiates BMP-induced smad signaling. Mol Cell 2013;51:5-19. ArticlePubMedPMC
- 32. Zhang L, Tran NT, Su H, Wang R, Lu Y, Tang H, Aoyagi S, Guo A, Khodadadi-Jamayran A, Zhou D, Qian K, Hricik T, Cote J, Han X, Zhou W, Laha S, Abdel-Wahab O, Levine RL, Raffel G, Liu Y, Chen D, Li H, Townes T, Wang H, Deng H, Zheng YG, Leslie C, Luo M, Zhao X. Cross-talk between PRMT1-mediated methylation and ubiquitylation on RBM15 controls RNA splicing. Elife 2015;4:e07938. ArticlePubMedPMCPDF
REFERENCES
Pancreas-specific protein arginine methyltransferase 1 (Prmt1) knock-out (KO) (Prmt1 PKO) mice exhibit a diabetic phenotype. (A) Representative images obtained by immunofluorescence (IF) staining of PRMT1 (green), pancreatic and duodenal homeobox 1 (PDX1, red), and 4′,6-diamidino-2-phenylindole (DAPI, blue) from wild type (WT) littermates and Prmt1 PKO mouse embryos at embryonic day 10.5 (E10.5). White scale bar, 50 µm. (B) Representative images of WT, Prmt1 heterozygous PKO (Het) littermates and Prmt1 PKO (KO) mice at postnatal day 7 (P7). (C) Body weights and (D) random blood glucose levels of WT, Prmt1 heterozygous PKO (Het) littermates and Prmt1 PKO (KO) mice at 4 weeks of age. (C, D) Each dot represents an individual data set from a given group. Lines and error bars indicate mean±standard error of mean (n=16 per group). aP<0.001 by one-way analysis of variance (ANOVA) with post hoc Bonferroni's test.
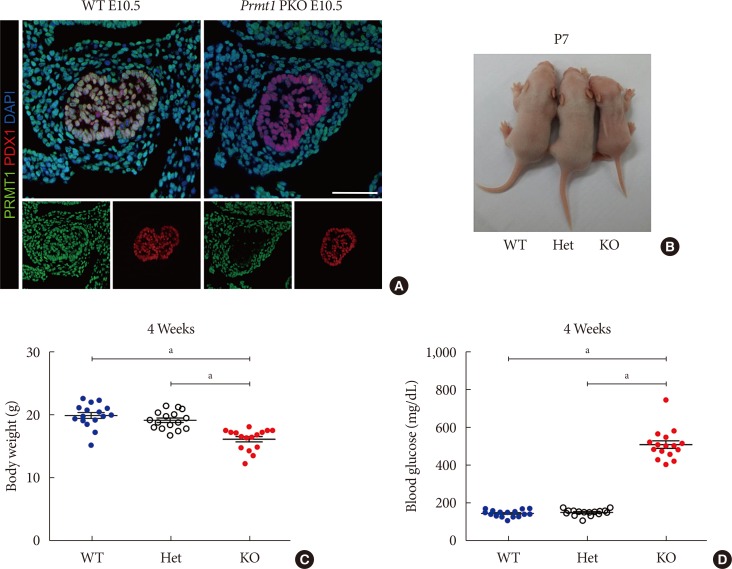
Pancreas-specific protein arginine methyltransferase 1 (Prmt1) knock-out (KO) (Prmt1 PKO) mice show hypoplastic pancreas with reduced β-cell mass. (A) Representative images of gastrointestinal tracts from wild type (WT) littermates and Prmt1 PKO mice at postnatal day 7 (P7). (B) Representative images obtained by hematoxylin and eosin (H&E) staining of pancreatic samples from WT littermates and Prmt1 PKO mice at P0 (black scale bar, 50 µm). (C) Representative images obtained by immunofluorescent staining of insulin (INS, green), amylase (AMY, red), mucin 1 (MUC1, white), and 4′,6-diamidino-2-phenylindole (DAPI, blue) from WT littermates and Prmt1 PKO mice at P0 (white scale bar, 50 µm). (D) Quantification of insulin-positive β-cell areas from WT littermates and Prmt1 PKO mice at P0. Each dot represents an individual data sets from a given group. Lines and error bars indicate mean±standard error of mean (n=4 per group). aP<0.001 by Student's t-test.
Protein arginine methyltransferase 1 (PRMT1) is required for endocrine cell commitment during the secondary transition. (A) Representative images obtained by immunofluorescent staining of SRY-box 9 (SOX9) or pancreatic and duodenal homeobox 1 (PDX1, green), NK2 homeobox 2 (NKX2.2), glucagon (GCG) or neurogenin 3 (NGN3, red), mucin 1 (MUC1, white), and 4′,6-diamidino-2-phenylindole (DAPI, blue) from wild type (WT) littermates and Prmt1 PKO mouse embryos at embryonic day 13.5 (E13.5) (white scale bar, 50 µm). (B) Representative images obtained by immunofluorescent staining of SOX9 (green), NKX2.2, NKX6.1 or ISL LIM homeobox 1 (ISL1, red) and DAPI (blue) from WT littermates and Prmt1 PKO mouse embryos at E14.5 (white scale bar, 50 µm).
Pancreas-specific protein arginine methyltransferase 1 (Prmt1) knock-out (KO) (Prmt1 PKO) mice exhibit increased numbers of neurogenin 3 (NGN3)-expressing cells during the secondary transition. (A) Representative images obtained by immunofluorescent staining of SRY-box 9 (SOX9, green), NGN3 (red), mucin 1 (MUC1, white) and 4′,6-diamidino-2-phenylindole (DAPI, blue) from wild type (WT) littermates and Prmt1 PKO mouse embryos at embryonic day 14.5 (E14.5) (white scale bars, 50 µm). (B) Representative images obtained by immunofluorescent staining of pancreatic and duodenal homeobox 1 (PDX1, green), phosphorylated histone H3 (PHH3, red) and MUC1 (white) from WT littermates and Prmt1 PKO mouse embryos at E14.5 (white scale bar, 50 µm). (C) Quantification of PHH3-positive proliferative cells relative to PDX1-positive pancreatic progenitor cells from WT littermates and Prmt1 PKO (KO) mouse embryos at E14.5. Each dot represents an individual data set from a given group. Lines and error bars indicate mean±standard error of mean (n=3 per group). aP<0.05 by Student's t-test.
Protein arginine methyltransferase 1 (PRMT1) is necessary for rapid degradation of the neurogenin 3 (NGN3) protein. (A) Representative images obtained by immunofluorescent staining of NGN3 (red) and mucin 1 (MUC1, white) from wild type (WT) littermates and Prmt1 PKO mouse embryos at embryonic day 14.5 (E14.5), E16.5, and E18.5 (white scale bar, 50 µm). (B) Relative Neurog3 expression levels were assessed by quantitative reverse transcription polymerase chain reaction of pancreatic samples from WT littermates and Prmt1 PKO mouse embryos at E13.5, E14.5, E16.5, E17.5, E18.5, and postnatal day 0 (P0). Data are expressed as the mean±standard error of mean (n=5 or 6 per group). (C) Immunoblot analysis detecting NGN3 protein levels in the pancreas of WT, Prmt1 heterozygous PKO littermates and Prmt1 PKO mouse embryos at E14.5. PRMT1 and α-tubulin were included in the analysis as reference proteins. (D) mPAC cells were transduced with adenovirus expressing nonspecific RNAi or Prmt1 RNAi and transfected with a vector encoding hemagglutinin (HA)-NGN3. Cells were then treated with cycloheximide (CHX, 100 µg/mL) for the indicated durations. NGN3 protein levels were determined by immunoblot analysis. α-Tubulin was included as a reference protein. KD, knockdown. aP<0.05, bP<0.01 by Student's t-test.
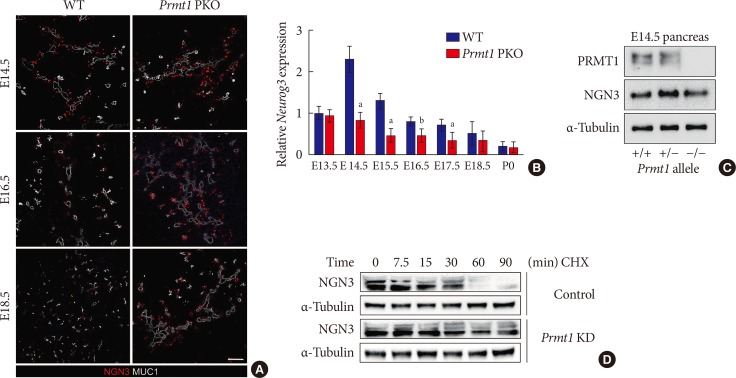
Figure & Data
References
Citations

- Arginine 65 methylation of Neurogenin 3 by PRMT1 is required for pancreatic endocrine development of hESCs
Gahyang Cho, Kwangbeom Hyun, Jieun Choi, Eunji Shin, Bumsoo Kim, Hail Kim, Jaehoon Kim, Yong-Mahn Han
Experimental & Molecular Medicine.2023; 55(7): 1506. CrossRef - Protein arginine methyltransferase 1 in the generation of immune megakaryocytes: A perspective review
Xinyang Zhao, Zechen Chong, Yabing Chen, X. Long Zheng, Qian-Fei Wang, Yueying Li
Journal of Biological Chemistry.2022; 298(11): 102517. CrossRef - Arginine 65 Methylation of Neurogenin 3 by PRMT1 Is Required for Pancreatic Endocrine Development of hESCs
Gahyang Cho, Kwangbeom Hyun, Jieun Choi, Eun Ji Shin, Bumsoo Kim, Hail Kim, Jaehoon Kim, Yong-Mahn Han
SSRN Electronic Journal .2022;[Epub] CrossRef - Protein Arginine Methyltransferase 1 Is Essential for the Meiosis of Male Germ Cells
Sahar Waseem, Sudeep Kumar, Kanghoon Lee, Byoung-Ha Yoon, Mirang Kim, Hail Kim, Keesook Lee
International Journal of Molecular Sciences.2021; 22(15): 7951. CrossRef - Proteome-Wide Alterations of Asymmetric Arginine Dimethylation Associated With Pancreatic Ductal Adenocarcinoma Pathogenesis
Meijin Wei, Chaochao Tan, Zhouqin Tang, Yingying Lian, Ying Huang, Yi Chen, Congwei Chen, Wen Zhou, Tao Cai, Jiliang Hu
Frontiers in Cell and Developmental Biology.2020;[Epub] CrossRef

 KDA
KDA PubReader
PubReader Cite
Cite