- Current
- Browse
- Collections
-
For contributors
- For Authors
- Instructions to authors
- Article processing charge
- e-submission
- For Reviewers
- Instructions for reviewers
- How to become a reviewer
- Best reviewers
- For Readers
- Readership
- Subscription
- Permission guidelines
- About
- Editorial policy
Articles
- Page Path
- HOME > Diabetes Metab J > Volume 39(6); 2015 > Article
-
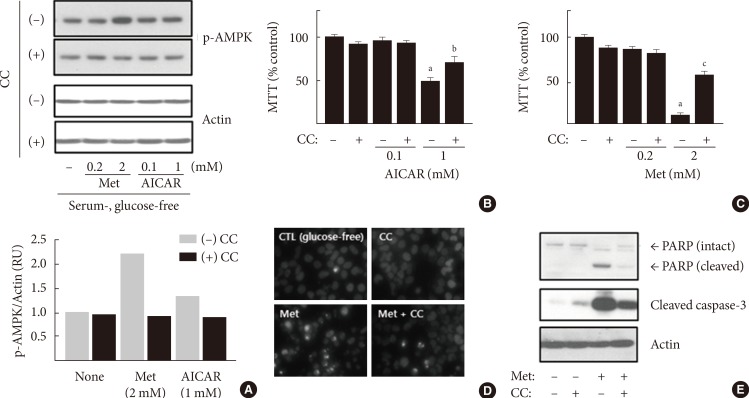
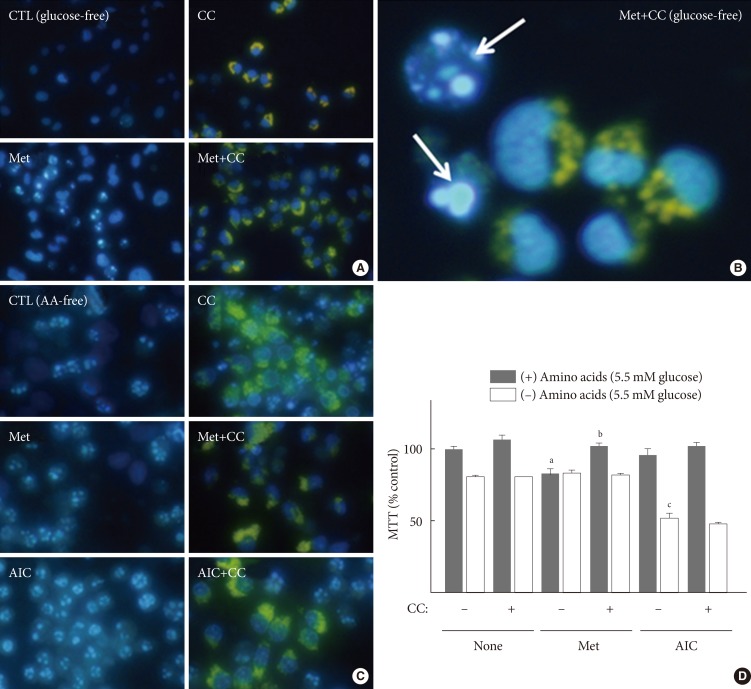
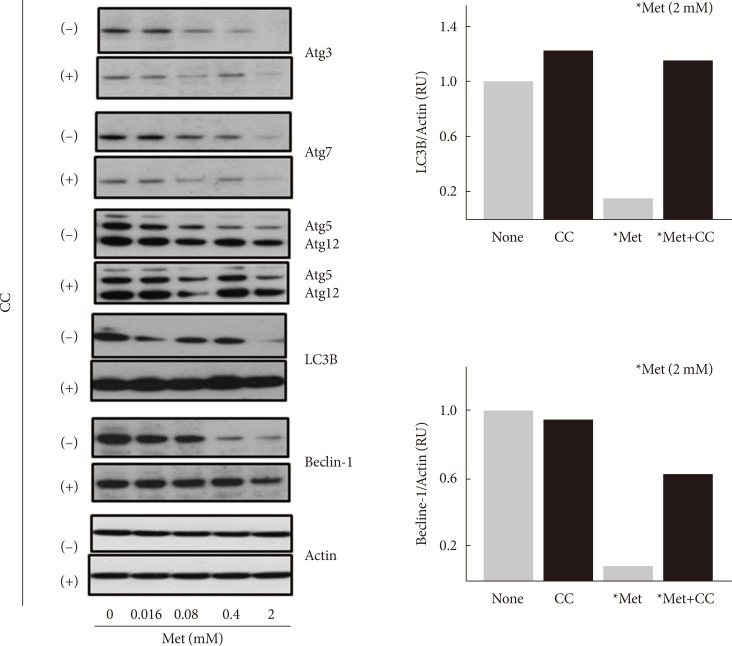
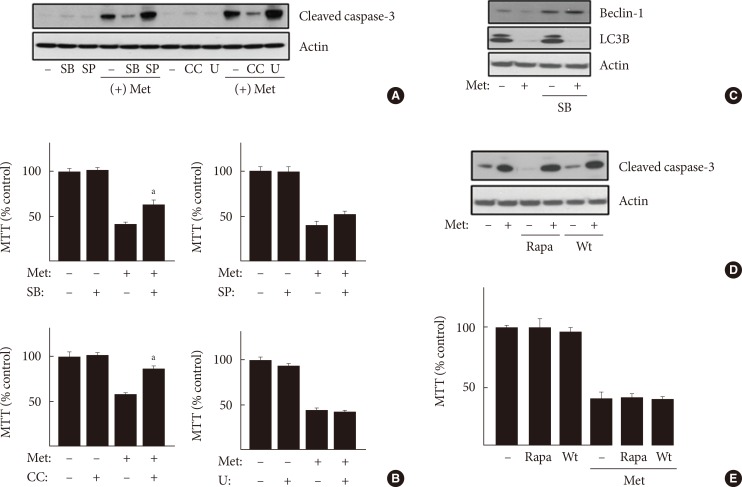
Original ArticleOthers Metformin Promotes Apoptosis but Suppresses Autophagy in Glucose-Deprived H4IIE Hepatocellular Carcinoma Cells
- Deok-Bae Park
-
Diabetes & Metabolism Journal 2015;39(6):518-527.
DOI: https://doi.org/10.4093/dmj.2015.39.6.518
Published online: December 11, 2015
Department of Histology and Institute of Medical Science, Jeju National University School of Medicine, Jeju, Korea.
- Corresponding author: Deok-Bae Park. Department of Histology and Institute of Medical Science, Jeju National University School of Medicine, 102 Jejudaehang-ro, Jeju 54987, Korea. parkdb@jejunu.ac.kr
• Received: November 17, 2014 • Accepted: February 26, 2015
Copyright © 2015 Korean Diabetes Association
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Figure & Data
References
Citations
Citations to this article as recorded by 

- Metformin Induces Lipogenesis and Apoptosis in H4IIE Hepatocellular
Carcinoma Cells
Deokbae Park, Sookyoung Lee, Hyejin Boo
Development & Reproduction.2023; 27(2): 77. CrossRef - Novel phloretin-based combinations targeting glucose metabolism in hepatocellular carcinoma through GLUT2/PEPCK axis of action: in silico molecular modelling and in vivo studies
Alaa Elmetwalli, Neamat H. Kamosh, Rania El Safty, Amany I. Youssef, Mohammed M. Salama, Khaled M. Abd El-Razek, Tarek El-Sewedy
Medical Oncology.2023;[Epub] CrossRef - Targeted Pyroptosis Is a Potential Therapeutic Strategy for Cancer
Hao Wu, Dianlun Qian, Xiangfeng Bai, Shibo Sun, Jayaprakash Narayana Kolla
Journal of Oncology.2022; 2022: 1. CrossRef - The effects of metformin on autophagy
Guangli Lu, Zhen Wu, Jia Shang, Zhenxing Xie, Chaoran Chen, Chuning zhang
Biomedicine & Pharmacotherapy.2021; 137: 111286. CrossRef - Protective Effect of Metformin against Hydrogen Peroxide-Induced Oxidative Damage in Human Retinal Pigment Epithelial (RPE) Cells by Enhancing Autophagy through Activation of AMPK Pathway
Xia Zhao, Linlin Liu, Yizhou Jiang, Marta Silva, Xuechu Zhen, Wenhua Zheng
Oxidative Medicine and Cellular Longevity.2020; 2020: 1. CrossRef Metformin Induces Autophagy via the AMPK-mTOR Signaling Pathway in Human Hepatocellular Carcinoma Cells
Chun Gao, Long Fang, Hui Zhang, Wei-Shuo Zhang, Xiao-Ou Li, Shi-Yu Du
Cancer Management and Research.2020; Volume 12: 5803. CrossRef- Metabolomics profiling of metformin-mediated metabolic reprogramming bypassing AMPKα
Min Yan, Huan Qi, Tian Xia, Xinjie Zhao, Wen Wang, Zhichao Wang, Chang Lu, Zhen Ning, Huan Chen, Tongming Li, Dinesh Singh Tekcham, Xiumei Liu, Jing Liu, Di Chen, Xiaolong Liu, Guowang Xu, Hai-long Piao
Metabolism.2019; 91: 18. CrossRef - Metformin Induces Oxidative Stress-Mediated Apoptosis without the Blockade of Glycolysis in H4IIE Hepatocellular Carcinoma Cells
Deokbae Park
Biological and Pharmaceutical Bulletin.2019; 42(12): 2002. CrossRef - Activation of AMPK prevents monocrotaline-induced pulmonary arterial hypertension by suppression of NF-κB-mediated autophagy activation
Cui Zhai, Wenhua Shi, Wei Feng, Yanting Zhu, Jian Wang, Shaojun Li, Xin Yan, Qingting Wang, Qianqian Zhang, Limin Chai, Cong Li, Pengtao Liu, Manxiang Li
Life Sciences.2018; 208: 87. CrossRef - Metformin and epothilone A treatment up regulate pro-apoptotic PARP-1, Casp-3 and H2AX genes and decrease of AKT kinase level to control cell death of human hepatocellular carcinoma and ovary adenocarcinoma cells
Aneta Rogalska, Barbara Bukowska, Agnieszka Marczak
Toxicology in Vitro.2018; 47: 48. CrossRef - Quantitative assessment of cell fate decision between autophagy and apoptosis
Bing Liu, Zoltán N. Oltvai, Hülya Bayır, Gary A. Silverman, Stephen C. Pak, David H. Perlmutter, Ivet Bahar
Scientific Reports.2017;[Epub] CrossRef - Meta-analysis of studies using metformin as a reducer for liver cancer risk in diabetic patients
Shujuan Ma, Yixiang Zheng, Yanni Xiao, Pengcheng Zhou, Hongzhuan Tan
Medicine.2017; 96(19): e6888. CrossRef - ROS Production and ERK Activity Are Involved in the Effects of d-β-Hydroxybutyrate and Metformin in a Glucose Deficient Condition
Santosh Lamichhane, Tonking Bastola, Ramesh Pariyar, Eun-Sol Lee, Ho-Sub Lee, Dae Lee, Jungwon Seo
International Journal of Molecular Sciences.2017; 18(3): 674. CrossRef - Metformin represses glucose starvation induced autophagic response in microvascular endothelial cells and promotes cell death
Samson Mathews Samuel, Suparna Ghosh, Yasser Majeed, Gnanapragasam Arunachalam, Mohamed M. Emara, Hong Ding, Chris R. Triggle
Biochemical Pharmacology.2017; 132: 118. CrossRef - NHX-5, an Endosomal Na+/H+ Exchanger, Is Associated with Metformin Action
Jeongho Kim, Hye-Yeon Lee, Jheesoo Ahn, Moonjung Hyun, Inhwan Lee, Kyung-Jin Min, Young-Jai You
Journal of Biological Chemistry.2016; 291(35): 18591. CrossRef - Metformin in pancreatic cancer treatment: from clinical trials through basic research to biomarker quantification
Archana Bhaw-Luximon, Dhanjay Jhurry
Journal of Cancer Research and Clinical Oncology.2016; 142(10): 2159. CrossRef - Metformina: stary lek w nowej aplikacji
Anna Dmoszyńska, Monika Podhorecka, Krzysztof Giannopoulos
Acta Haematologica Polonica.2016; 47(2): 139. CrossRef

 KDA
KDA

 PubReader
PubReader Cite
Cite