R1467H Variants of Rho Guanine Nucleotide Exchange Factor 11 (ARHGEF11) are Associated with Type 2 Diabetes Mellitus in Koreans
Article information
Abstract
Background
The human Rho guanine nucleotide exchange factor 11 (ARHGEF11) functions as an activator of Rho GTPases and is thought to influence insulin signaling. The R1467H variant of ARHGEF11 has been reported to be associated with susceptibility to type 2 diabetes mellitus (T2DM) in Western populations.
Methods
We investigated the effects of the R1467H variant on susceptibility to T2DM as well as related traits in a Korean population. We genotyped the R1467H (rs945508) of ARHGEF11 in 689 unrelated T2DM patients and 249 non-diabetic individuals and compared the clinical and biochemical characteristics according to different alleles.
Results
The H allele was significantly more frequent in T2DM cases than in controls (P = 0.037, 17.1% and 13.1%; respectively). H homozygocity was associated with a higher risk of T2DM compared to those with R/R or R/H genotype (odds ratio, 5.24; 95% confidence interval, 1.06 to 25.83; P = 0.042). The fasting plasma glucose, HbA1c, fasting insulin, HOMA2-IR and HOMA2-%β levels did not differ significantly between different genotypes.
Conclusion
Our study replicated associations of the ARHGEF11 polymorphism with increased risk of T2DM in a Korean population and thus supports previous data implicating a potential role of ARHGEF11 in the etiology of T2DM. Further studies revealing the underlying mechanism for this association are needed.
INTRODUCTION
Type 2 diabetes mellitus (T2DM) is a multi-factorial trait, in which individual risk is defined by the complex interplay of genetic and environmental factors [1-3]. To search for genetic loci that contribute to T2DM, several genomic linkage scans have been completed worldwide [4,5]. Several previous studies have reported evidence for linkage between T2DM and the chromosome 1q21-q24 region [6-8] and have also reported that this region contributes to impaired glucose tolerance (IGT) [9]. In the Pima Indians and Amish populations, investigators genotyped multiple single nucleotide polymorphisms (SNPs) within this region, and results pointed to a region including the Rho guanine nucleotide exchange factor 11 (ARHGEF11) gene [10,11]. ARHGEF11 is an activator of Rho GTPases and may play a role in the G protein signaling involved in β-cell apoptosis, insulin secretion and insulin signaling. ARHGEF11 is expressed in the pancreas, liver, muscle, and adipose tissue [12-14]. Previous studies have implicated Rho subfamily G proteins in physiological insulin secretion [13,14], and Rho GT-Pases have been suggested to play a role in insulin signaling through the activation of p38 mitogen activated protein kinase and Jun NH2-terminal kinase pathways [15-17]. Moreover, Rho family small GTP-binding protein TC10 activation and phosphatidylinositol 3-kinase activation regulates the dynamic actin rearrangement required for insulin-stimulated translocation of GLUT4 [18,19]. Together, these data suggest that proteins involved in G protein signaling, such as ARHGEF11, may play an important role in glucose homeostasis.
The full-length ARHGEF11 protein contains 1,522 amino acids, has a calculated molecular mass of 168.6 kD and includes 41 exons [20]. The C-terminal (amino acids 1,181-1,522) region of ARHGEF11 protein has been shown to regulate ARHGEF11 protein activity [21]. Recently, a non-synonymous amino acid substitution variant within the C-terminal region of ARHGEF11 R1467H (rs945508, G/A), was reported to be associated with T2DM in Western sample populations [11,22,23]. In the Amish, the 1467R allele was the risk allele for T2DM, while the 1467H allele was the risk allele for insulin resistance and T2DM in Pima Indians [11,22]. Moreover, replicated association of the 1467H allele as the risk allele for T2DM and/or IGT in a German Caucasian population have been reported [23]. This inconsistency in the effects of genetic variants may be due to different genetic backgrounds between the different ethnic groups. Therefore, in the present study, we evaluated the effects of this variant on the susceptibility to T2DM as well as its relation to metabolic parameters in a Korean population.
METHODS
Study subjects
A total of 689 unrelated T2DM patients and 249 non-diabetic individuals were recruited at the Inha University Hospital of Korea from April 2004 to December 2007. Diabetes was defined according to the 1997 American Diabetes Association diagnostic criteria [24]. Exclusion criteria were type 1 diabetes mellitus, current malignancy, history of ketoacidosis, positive for glutamic acid decarboxylase antibody, severe kidney or liver disease, and corticosteroid use. All subjects gave written informed consent before taking part in the study, which was approved by the Inha University Hospital Institutional Review Board.
Single nucleotide polymorphism genotyping
Genomic DNA was extracted from leukocytes in the whole-blood samples using a Gentra Puregene Blood Kit Plus (Qiagen, Valencia, CA, USA). Genotyping of the R1467H was achieved using the TaqMan allelic discrimination assay (Applied Biosystems, Foster City, CA, USA). To assess genotyping reproducibility, a random -10% selection of the samples was re-genotyped in all SNPs with 100% concordance. The SNPs were in accordance with Hardy-Weinberg equilibrium in the T2DM subjects (P > 0.10) and in the control subjects (P > 0.20).
Clinical and biochemical measurements
Baseline laboratory data and anthropometric measurements were collected for each patient. Low density lipoprotein cholesterol levels were calculated using the Friedewald formula [25], and plasma glucose was measured using a glucose oxidase method. Hemoglobin A1c (HbA1c) values were determined using high-performance liquid chromatography, and insulin concentrations were measured using a radioimmunoassay kit (IRMA kit; DAINABOT, Tokyo, Japan). The insulin resistance and insulin secretion indexes were calculated using the HOMA2 model (HOMA2-IR and HOMA2-%β, respectively), i.e., the updated computer model for the pairing of fasting plasma glucose and insulin (The HOMA2 model is available from www.ocdem.ox.ac.uk) [26].
Statistical analysis
Statistical analyses were performed using SPSS for Windows version 18.0 (SPSS Inc., Chicago, IL, USA), and data are shown as means ± standard deviation, unless otherwise indicated. Associations between categorical variables were analyzed using χ2 tests and continuous variables were assessed using Student's t-test. The differences in allelic and genotypic frequencies between the diabetic and control individuals were analyzed using Pearson's χ2 test. Fasting insulin, HOMA2-IR and HOMA2-%β variables were log-transformed to correct for skew. The odds ratio (OR) was assessed by counting the number of risk alleles for each individual, and association of genotypes with T2DM was assessed via a logistic regression analysis adjusted for age, gender, and body mass index (BMI). All P < 0.05 were considered to be significant.
RESULTS
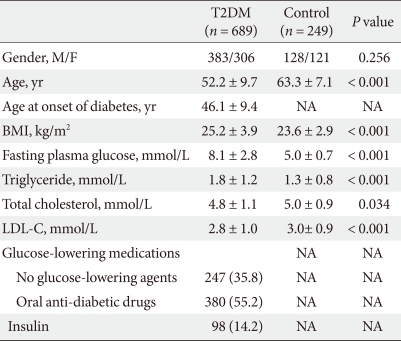
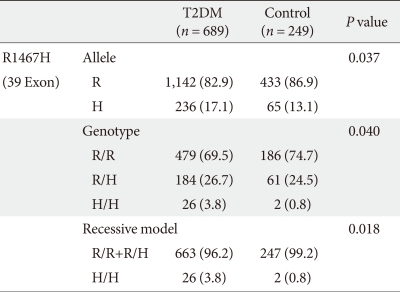
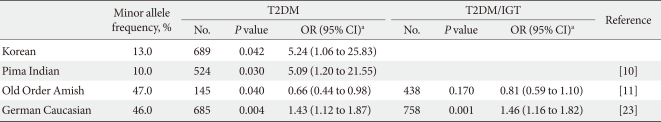
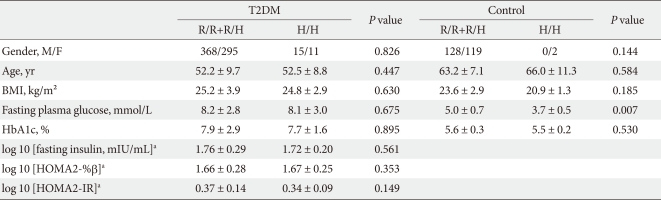
We performed a case-control study including 249 non-diabetic individuals and 689 unrelated T2DM patients in Korea. The group of patients with T2DM included 383 men and 306 women, and the control group included 128 men and 121 women (P = 0.256). Non-diabetic individuals were older than T2DM patients (P < 0.001, 63.25 ± 7.12, 52.23 ± 9.70 years, respectively) and had a lower BMI 23.59 ± 2.89 kg/m2 (P < 0.001, 25.21 ± 3.87 kg/m2). The clinical features of the participants are summarized in Table 1. Allelic and genotypic frequencies of the SNP in ARHGEF11 are shown in Table 2. The H allele was significantly more frequent in T2DM cases than in controls (17.1% in T2DM vs. 13.1% in control group, P = 0.037). Genotypic frequencies were significantly different in T2DM patients compared to those of the controls (P = 0.040, R/R 479 [69.5%] vs. 186 [74.7%], R/H 184 [26.7%] vs. 61 [24.5%], H/H 26 [3.8%] vs. 2 [0.8%], respectively). Moreover, in patients with T2DM, the probability of having the H/H genotype vs. R/R + H/H was significantly higher compared to that of the control subjects (P = 0.018, 3.8% vs. 0.8%), and this association was significant after adjustment for age, gender and BMI (OR, 5.24; 95% confidence intervals [CI], 1.06-25.83; P = 0.042). The odds ratio estimates along with 95% CI values for the ARHGEF11 R1467H allele in various studies conducted in T2DM patients are presented in Table 3. In order to further examine the role of the H allele in T2DM subjects, we assessed whether the H homozygote was associated with insulin secretion or insulin resistance in T2DM subjects. There was no significant difference between R/R or R/H subjects and H/H subjects with regard to fasting plasma glucose, HbA1c, fasting insulin, HOMA2-IR or HOMA2-%β (P > 0.05, Table 4).
DISCUSSION
In this study, we found a significant association between the variation at R1467H in ARHGEF11 and T2DM in a Korean population, similar to the results previously reported in Pima Indians and German Caucasians [10,23]. It is noteworthy that the allele frequency and odds ratio in our study are closer to those in Pima Indians (Table 3) and that the association between the variation at R1467H in ARHGEF11 and T2DM which has been observed in other ethnic groups has been replicated, supporting the reliabilities of both previous studies as well as that of our present study. However, the association with type 2 diabetes may have a stronger impact in the German Caucasians than in Koreans due to the higher frequency of the R1467H risk allele (H) in German Caucasians (46% vs. 13%). The lower allelic frequency in Koreans is similar to the data reported by the HapMap project [27]. A different association of the common R allele with T2DM was shown in a study of the Amish [11]. This may be due to population differences; however, considering the Swiss-German ancestry of the Amish, it was surprising that data from German Caucasians more closely adhered to the Pima findings rather than those reported in the Amish. Although, there is no evidence that the R1467H variant may exert an effect on ARHGEF11 expression, an Arg to His amino acid change in the C-terminal region of ARHGEF11 may be brought about by the variant. Thus, we believe that our present findings are consistent with those found in Pima Indians and German Caucasians, strongly suggesting the role of the variant in the pathophysiology of T2DM.
The H allele at R1467H in ARHGEF11 increases the risk for T2DM in Pima Indians by reducing insulin-stimulated glucose uptake. Among the non-diabetic, full-heritage Pima Indians, subjects with an H allele had a lower mean glucose disposal rate during a hyperinsulinemic-euglycemic clamp. To further illuminate the role of the H allele in a population of Korean T2DM patients, related traits were compared in subjects with H/H homozygocity and in those carrying the major R allele (R/H or R/R). Our study did not reveal any significant differences in fasting plasma glucose, HbA1c, fasting insulin, HOMA2-IR and HOMA2-%β levels in T2DM subjects between the two groups. The H allele also did not increase fasting plasma glucose in the control patients, possibly due to the various drug effects experienced by T2DM patients or that the HOMA parameters are a relatively crude analysis method compared to the clamp method [28]. It may also be due to different ethnic and environmental backgrounds. Therefore, additional investigations in independent populations, as well as functional studies, will be necessary to clarify the influence of the ARHGEF11 variant on T2DM. There is also a possibility that the R1467H variant may not be the true functional variant but rather a variant in linkage disequilibrium with the true functional variant.
This study had several limitations. First, our study was similar to others in Pima Indians and had similar associations between the variations at R1467H in ARHGEF11 and T2DM but could not determine a significant association between the R/R+R/H and H/H groups of T2DM with regard to fasting plasma glucose, HbA1c, fasting insulin, HOMA2-IR or HOMA2-%β level. In Pima Indians, the investigator demonstrated that the H allele at R1467H in ARHGEF11 increases the risk of T2DM by reducing insulin-stimulated glucose uptake in non-diabetic controls. However, our study did not measure insulin levels in non-diabetic control subjects. Second, we used fasting insulin concentration and the HOMA2 model as markers of insulin resistance. The HOMA2 model has been significantly correlated with the hyperinsulinemic-euglycemic clamp, which is accepted as the standard in defining insulin resistance; however, the HOMA2 model is described for approximate estimation [28]. Finally, because our study was conducted in a Korean population, we cannot generalize these findings to other ethnic groups.
In conclusion, our study replicated associations of the ARHGEF11 polymorphism with increased risk of T2DM in a Korean population and thus supports previous data implicating a potential role of ARHGEF11 in the etiology of T2DM. Further replication studies in different ethnic populations are necessary to confirm the effect of the R1467H variant on T2DM.
ACKNOWLEDGEMENT
This study was supported by a grant of the Korean Health 21 Research and Development Project, Ministry of Health & Welfare, Republic of Korea (A050463, A070001).